Knowledge graph surpassing 1 billion biomedical relations and a new Data Resources framework
Greetings and Happy New Year! We are happy to report that the Data4Cure Biomedical Intelligence® Cloud and CURIETM Knowledge Graph are once again significantly expanding to help with a yet broader set of tasks in pharmaceutical R&D.
With the updates this month, the CURIE Knowledge Graph is reaching 1 billion data-driven and literature-mined relations.
And, we are introducing a new way to semantically organize and keep track of thousands of public and internal data on the platform – the Data Resourcesframework.
Here is a quick break down of the platform by numbers:
State of the Biomedical Intelligence® Cloud:
Number of data tables (omics, phenotypic, clinical): ~100,000
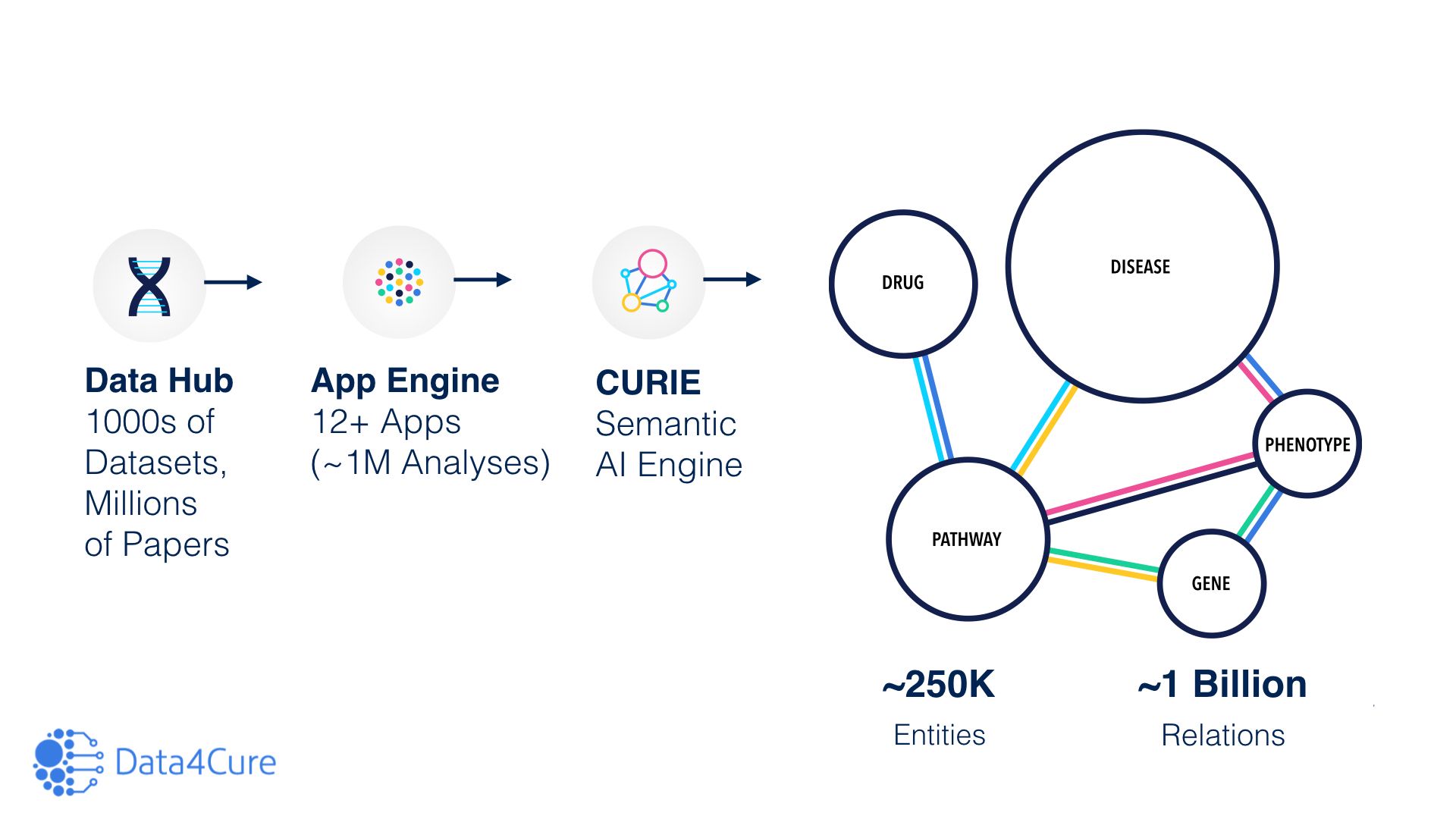
Number of analyses: ~1 million
Number of biomedical entities tracked: ~250,000
Number of relations in the CURIE Knowledge Graph: ~1 billion

Data Resources: a new way to thematically organize public and internal data in DataHub
The growth of the CURIE Knowledge Graph is facilitated by the continuous integration of new public data resources (and your internal resources for private Biomedical Intelligence Clouds).
Among the public data resources added most recently are data from NIH LINCS, Project DRIVE, new cell line omics data from DepMap, curated single-cell RNAseq data, new multi-omics, pathologic and clinical datasets from recentimmunotherapy clinical trials, and hundreds of curated datasets across inflammatory bowel disease (ulcerative colitis, Crohn’s disease), rheumatoid arthritis, non-alcoholic fatty liver disease and multiple myeloma. With these additions, there are now roughly 100,000 data tables in the system from public data alone.
To facilitate the integration but also fast retrieval, search and browsing of such a large and diverse collection of data, we are introducing a new component of the platform – the Data Resources framework which is now an integral part of the Data Hub application.
Data Resources take a project- or resource-centered view to data curation and presentation. Each Data Resource contains project metadata, project-associated data files, as well as analyses performed on the data. Data Resources can be accessed from the Data Hub application or directly from a CURIE Entity Page where resources specific to the biomedical entity of interest are organized and presented:

Resource metadata is fully searchable, semantically tagged & cross-linked via internal and external IDs. Resources are linked to underlying publications (if available) and can have associated lab notebook IDs or other internal or external identifiers. Resource data files and analyses are integrated with the CURIE Knowledge Graph, providing the ultimate framework for semantically linked omics, phenotypic, pathologic & clinical data. The Resource framework also includes fine-grained access control and sharing and the ability to connect internal and external data resources, extending on the capabilities previously available in the Data Hub.
The Data Resources framework is a new and powerful tool to help you move beyond analyzing one dataset at a time to leveraging thousands of semantically-linked datasets available on the platform to test hypotheses or gain new data-driven insights into a disease of interest. We will provide case studies with recent data resources in future posts in the weeks to come.
Now let’s turn data into cures.
We hope you found this update useful. As always, we’d love to hear from you. Write to us at info@data4cure.com. Let us know how we can help and let’s turn data into cures!
Until next time,
— The Data4Cure Team.